HMM Classifier¶
Multiple HMMs can be combined to form a multi-class classifier.
The HMMClassifier can be used to classify:
Univariate/multivariate numerical observation sequences, by using
GaussianMixtureHMMmodels.Univariate categorical observation sequences, by using
CategoricalHMMmodels.
To classify a new observation sequence \(O'\), the HMMClassifier works by:
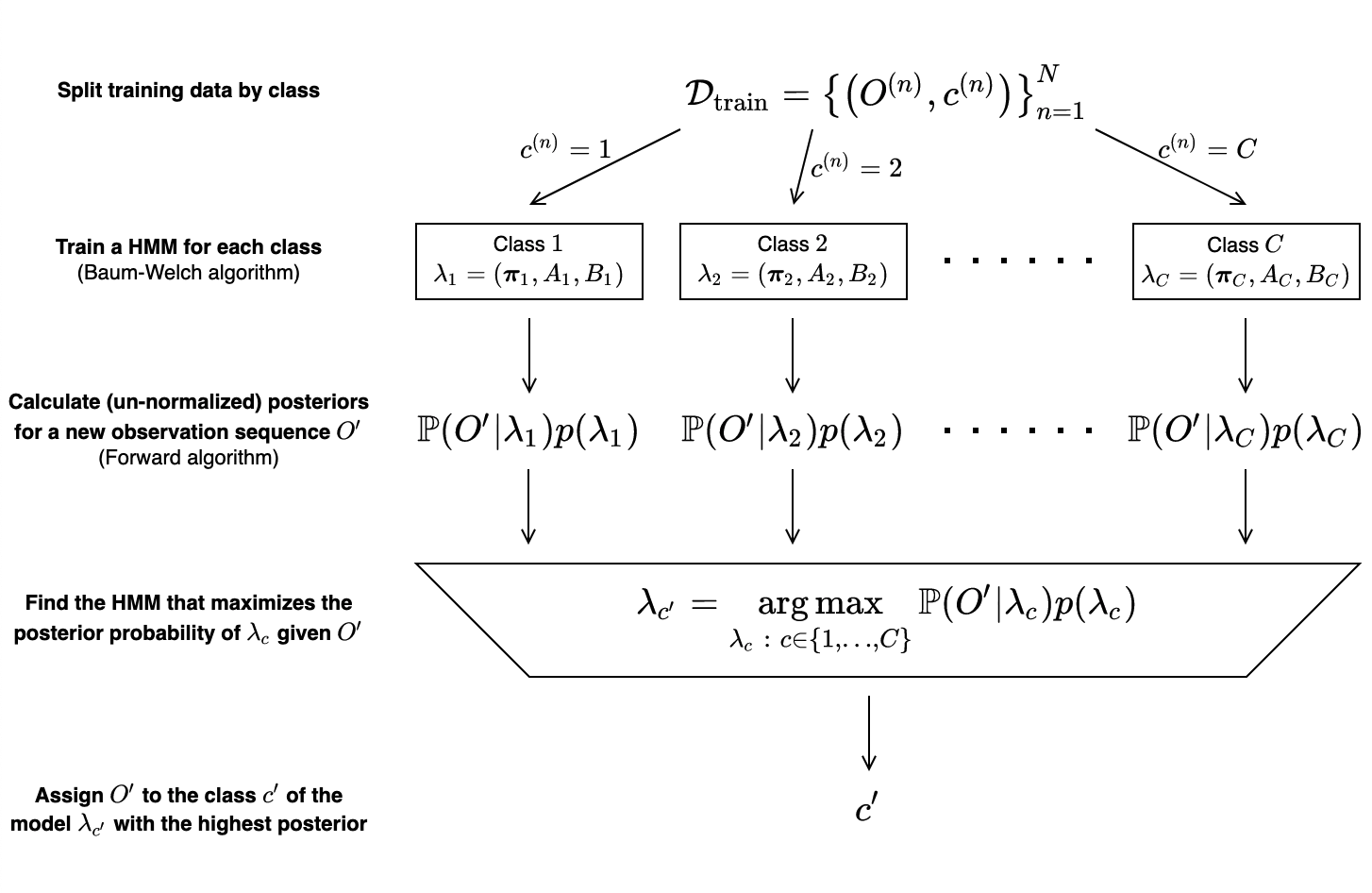
- Creating HMMs \(\lambda_1, \lambda_2, \ldots, \lambda_C\) and training each model using the Baum—Welch algorithm on the subset of training observation sequences with the same class label as the model.
- Calculating the likelihoods \(\mathbb{P}(O'\ |\ \lambda_1), \mathbb{P}(O'\ |\ \lambda_2), \ldots, \mathbb{P}(O'\ |\ \lambda_C)\) of each model generating \(O'\) using the Forward algorithm.
- Scaling the likelihoods by priors \(p(\lambda_1), p(\lambda_2), \ldots, p(\lambda_C)\), producing un-normalized posteriors\(\mathbb{P}(O'\ |\ \lambda_c)p(\lambda_c)\) for each class.
- Choosing the class represented by the HMM with the highest posterior probability for \(O'\).\[\begin{split}c' = \operatorname*{\arg\max}_{c\in\{1,\ldots,C\}}\ p(\lambda_c\ |\ O') = \operatorname*{\arg\\max}_{c\in\{1,\ldots,C\}}\ \mathbb{P}(O'\ |\ \lambda_c)p(\lambda_c)\end{split}\]
These steps are summarized in the diagram below.
API reference¶
Class¶
A classifier consisting of HMMs, each trained independently to recognize sequences of a single class. |
Methods¶
|
Initialize a |
|
Add a single HMM to the classifier. |
|
Add HMMs to the classifier. |
|
Fit the HMMs to the sequence(s) in |
|
Fit the model to the sequence(s) in |
|
Load and deserialize a fitted HMM classifier. |
|
Predict classes for the sequence(s) in |
|
Predict log un-normalized posterior probabilities for the sequences in |
|
Predict class probabilities for the sequence(s) in |
|
Predict class scores for the sequence(s) in |
|
Serialize and save a fitted HMM classifier. |
|
Calculate the predictive accuracy for the sequence(s) in |
- class sequentia.models.hmm.classifier.HMMClassifier¶
A classifier consisting of HMMs, each trained independently to recognize sequences of a single class.
The predicted class for a given observation sequence is the class represented by the HMM which produces the maximum posterior probability for the observation sequence.
Examples
Using a
HMMClassifier(withGaussianMixtureHMMmodels) to classify spoken digits.import numpy as np from sequentia.datasets import load_digits from sequentia.models.hmm import GaussianMixtureHMM, HMMClassifier # Seed for reproducible pseudo-randomness random_state = np.random.RandomState(1) # Fetch MFCCs of spoken digits data = load_digits() train_data, test_data = data.split(test_size=0.2, random_state=random_state) # Create a HMMClassifier using a class frequency prior clf = HMMClassifier(prior='frequency') # Add an untrained HMM for each class for label in data.classes: model = GaussianMixtureHMM(random_state=random_state) clf.add_model(model, label=label) # Fit the HMMs by providing training observation sequences for all classes clf.fit(train_data.X, train_data.y, lengths=train_data.lengths) # Predict classes for the test observation sequences y_pred = clf.predict(test_data.X, lengths=test_data.lengths)
As done in the above example, we can provide unfitted HMMs using
add_model()oradd_models(), then provide training observation sequences for all classes tofit(), which will automatically train each HMM on the appropriate subset of data.Alternatively, we may provide pre-fitted HMMs and call
fit()with no arguments.# Create a HMMClassifier using a class frequency prior clf = HMMClassifier(prior='frequency') # Manually fit each HMM on its own subset of data for X_train, lengths_train, label for train_data.iter_by_class(): model = GaussianMixtureHMM(random_state=random_state) model.fit(X_train, lengths=lengths_train) clf.add_model(model, label=label) # Fit the classifier clf.fit()
- __init__(*, prior=PriorMode.UNIFORM, classes=None, n_jobs=1)¶
Initialize a
HMMClassifier.- Parameters:
self (HMMClassifier) –
prior (PriorMode | dict[int, float]) –
Type of prior probability to assign to each HMM.
If
None, a uniform prior will be used, making each HMM equally likely.If
"frequency", the prior probability of each HMM is equal to the fraction of total observation sequences that the HMM was fitted with.If a
dict, custom prior probabilities can be assigned to each HMM. The keys should be the label of the class represented by the HMM, and the value should be the prior probability for the HMM.
Set of possible class labels.
If not provided, these will be determined from the training data labels.
If provided, output from methods such as
predict_proba()andpredict_scores()will follow the ordering of the class labels provided here.
n_jobs (int) –
Maximum number of concurrently running workers.
If 1, no parallelism is used at all (useful for debugging).
If -1, all CPUs are used.
If < -1,
(n_cpus + 1 + n_jobs)are used — e.g.n_jobs=-2uses all but one.
- Return type:
- add_model(model, /, *, label)¶
Add a single HMM to the classifier.
- Parameters:
self (HMMClassifier) –
model (BaseHMM) – HMM to add to the classifier.
label (int) – Class represented by the HMM.
- Returns:
The classifier.
- Return type:
Notes
All models added to the classifier must be of the same type — either
GaussianMixtureHMMorCategoricalHMM.
- add_models(models, /)¶
Add HMMs to the classifier.
- Parameters:
self (HMMClassifier) –
models (dict[int, BaseHMM]) – HMMs to add to the classifier. The key for each HMM should be the label of the class represented by the HMM.
- Returns:
The classifier.
- Return type:
Notes
All models added to the classifier must be of the same type — either
GaussianMixtureHMMorCategoricalHMM.
- fit(X=None, y=None, *, lengths=None)¶
Fit the HMMs to the sequence(s) in
X.If fitted models were provided with
add_model()oradd_models(), no arguments should be passed tofit().If unfitted models were provided with
add_model()oradd_models(), training dataX,yandlengthsmust be provided tofit().
- Parameters:
self (HMMClassifier) –
X (ndarray[Any, dtype[float64]] | ndarray[Any, dtype[int64]] | None) – Sequence(s).
y (ndarray[Any, dtype[int64]] | None) – Classes corresponding to sequence(s) in
X.lengths (ndarray[Any, dtype[int64]] | None) –
Lengths of the sequence(s) provided in
X.If
None, thenXis assumed to be a single sequence.len(X)should be equal tosum(lengths).
- Returns:
The fitted classifier
- Return type:
- fit_predict(X, y, *, lengths=None)¶
Fit the model to the sequence(s) in
Xand predicts outputs forX.- Parameters:
self (ClassifierMixin) –
X (ndarray[Any, dtype[float64]] | ndarray[Any, dtype[int64]]) – Sequence(s).
y (ndarray[Any, dtype[int64]]) – Outputs corresponding to sequence(s) in
X.lengths (ndarray[Any, dtype[int64]] | None) –
Lengths of the sequence(s) provided in
X.If
None, thenXis assumed to be a single sequence.len(X)should be equal tosum(lengths).
- Returns:
Output predictions.
- Return type:
- classmethod load(path, /)¶
Load and deserialize a fitted HMM classifier.
- Parameters:
cls (type[HMMClassifier]) –
path (str | Path | IO) – Location to load the serialized classifier from.
- Returns:
Fitted HMM classifier.
- Return type:
See also
saveSerialize and save a fitted HMM classifier.
- predict(X, *, lengths=None)¶
Predict classes for the sequence(s) in
X.- Parameters:
- Returns:
Class predictions.
- Return type:
Notes
This method requires a trained classifier — see
fit().
- predict_log_proba(X, *, lengths=None)¶
Predict log un-normalized posterior probabilities for the sequences in
X.- Parameters:
- Returns:
Log probabilities.
- Return type:
Notes
This method requires a trained classifier — see
fit().
- predict_proba(X, *, lengths=None)¶
Predict class probabilities for the sequence(s) in
X.Probabilities are calculated as the posterior probability of each HMM generating the sequence.
- Parameters:
- Returns:
Class membership probabilities.
- Return type:
Notes
This method requires a trained classifier — see
fit().
- predict_scores(X, *, lengths=None)¶
Predict class scores for the sequence(s) in
X.Scores are calculated as the log posterior probability of each HMM generating the sequence.
- Parameters:
- Returns:
Class scores.
- Return type:
Notes
This method requires a trained classifier — see
fit().
- save(path, /)¶
Serialize and save a fitted HMM classifier.
- Parameters:
self (HMMClassifier) –
path (str | Path | IO) – Location to save the serialized classifier.
- Return type:
None
Notes
This method requires a trained classifier — see
fit().See also
loadLoad and deserialize a fitted HMM classifier.
- score(X, y, *, lengths=None, normalize=True, sample_weight=None)¶
Calculate the predictive accuracy for the sequence(s) in
X.- Parameters:
self (ClassifierMixin) –
X (ndarray[Any, dtype[float64]] | ndarray[Any, dtype[int64]]) – Sequence(s).
y (ndarray[Any, dtype[int64]]) – Outputs corresponding to sequence(s) in
X.lengths (ndarray[Any, dtype[int64]] | None) –
Lengths of the sequence(s) provided in
X.If
None, thenXis assumed to be a single sequence.len(X)should be equal tosum(lengths).
normalize (bool) – See
sklearn.metrics.accuracy_score().sample_weight (ndarray[Any, dtype[float64]] | ndarray[Any, dtype[int64]] | None) – See
sklearn.metrics.accuracy_score().
- Returns:
Predictive accuracy.
- Return type:
Notes
This method requires a trained classifier — see
fit().
- models: dict[int, BaseHMM]¶
HMMs constituting the
HMMClassifier.