Gaussian Mixture HMM¶
The Gaussian Mixture HMM is a variant of HMM that uses a multivariate Gaussian mixture model as the emission distribution for each state.
This HMM variant can be used to recognize unbounded real-valued univariate or multivariate sequences.
Emissions¶
The emission distribution \(b_m\) of an observation \(\mathbf{o}^{(t)}\) at time \(t\) for state \(m\) is formed by a weighted mixture of \(K\) multivariate Gaussian probability density functions, defined as:
Where:
- \(\mathbf{o}^{(t)}=\left(o_1^{(t)}, o_2^{(t)}, \ldots, o_D^{(t)}\right)\) is a single observation at time \(t\), such that \(\mathbf{o}^{(t)}\in\mathbb{R}^D\).
- \(c_k^{(m)}\) is a component mixture weight for the \(k^\text{th}\) Gaussian component of the \(m^\text{th}\) state, such that \(\sum_{k=1}^K c_k^{(m)} = 1\) and \(c_k^{(m)}\in[0, 1]\).
- \(\boldsymbol\mu_k^{(m)}\) is a mean vector for the \(k^\text{th}\) Gaussian component of the \(m^\text{th}\) state, such that \(\boldsymbol\mu_k^{(m)}\in\mathbb{R}^D\).
- \(\Sigma_k^{(m)}\) is a covariance matrix for the \(k^\text{th}\) Gaussian component of the \(m^\text{th}\) state, such that \(\Sigma_k^{(m)}\in\mathbb{R}^{D\times D}\) and \(\Sigma_k^{(m)}\) is symmetric and positive semi-definite.
- \(\mathcal{N}_D\) is the \(D\)-dimensional multivariate Gaussian probability density function.
Using a mixture rather than a single Gaussian allows for more flexible modelling of observations.
The component mixture weights, mean vector and covariance matrix for all states and Gaussian components are updated during training via Expectation-Maximization through the Baum-Welch algorithm.
Covariance matrix types¶
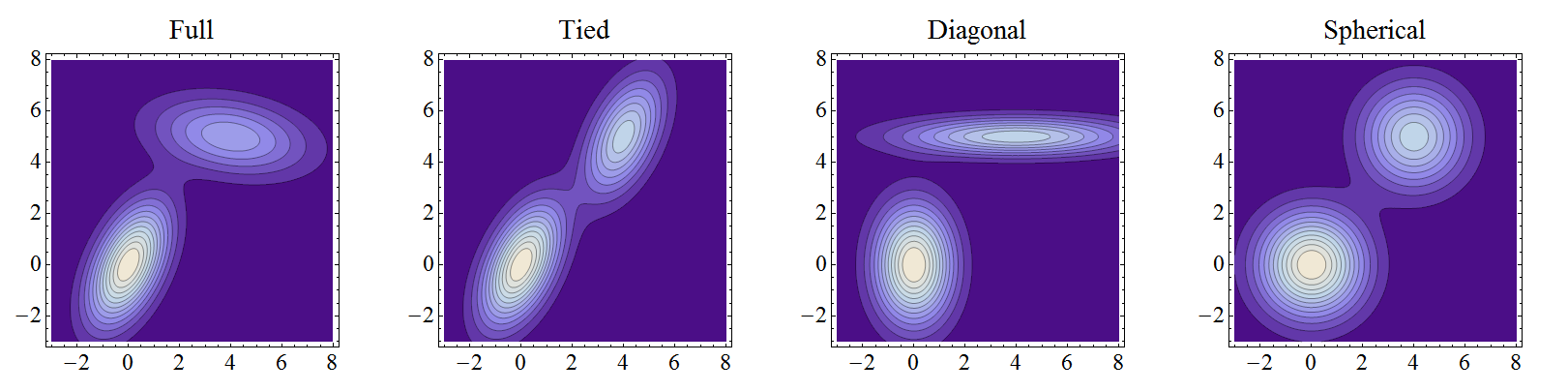
The \(K\) covariance matrices for a state can come in different forms:
Full: All values are fully learnable independently for each component.
Diagonal: Only values along the diagonal may be learned independently for each component.
Spherical: Same as diagonal, with a single value shared along the diagonal for each component.
Tied: Same as full, with all components sharing the same single covariance matrix.
Estimating a full covariance matrix is not always necessary, particularly when a sufficient number of Gaussian components are used. If time is limiting, a spherical, diagonal and tied covariance matrix may also yield strong results while reducing training time due to having fewer parameters to estimate.
API reference¶
Class¶
A hidden Markov model with multivariate Gaussian mixture emissions. |
Methods¶
|
Initializes the |
|
The Akaike information criterion of the model, evaluated with the maximum likelihood of |
|
The Bayesian information criterion of the model, evaluated with the maximum likelihood of |
|
Fit the HMM to the sequences in |
|
Freeze the trainable parameters of the HMM, preventing them from be updated during the Baum—Welch algorithm. |
|
Calculate the log-likelihood of the HMM generating a single observation sequence. |
|
Set the covariance matrices of the state emission distributions. |
|
Set the mean vectors of the state emission distributions. |
|
Set the initial state probabilities. |
|
Set the transition probability matrix. |
|
Set the component mixture weights of the state emission distributions. |
|
Unfreeze the trainable parameters of the HMM, allowing them to be updated during the Baum—Welch algorithm. |
Number of trainable parameters — requires |
- class sequentia.models.hmm.variants.GaussianMixtureHMM¶
A hidden Markov model with multivariate Gaussian mixture emissions.
Examples
Using a
GaussianMixtureHMMto learn how to recognize spoken samples of the digit 3.See
load_digits()for more information on the sample dataset used in this example.import numpy as np from sequentia.datasets import load_digits from sequentia.models.hmm import GaussianMixtureHMM # Seed for reproducible pseudo-randomness random_state = np.random.RandomState(1) # Fetch MFCCs of spoken samples for the digit 3 data = load_digits(digits=[3]) train_data, test_data = data.split(test_size=0.2, random_state=random_state) # Create and train a GaussianMixtureHMM to recognize the digit 3 model = GaussianMixtureHMM(random_state=random_state) X_train, lengths_train = train_data.X_lengths model.fit(train_data.X, lengths=train_data.lengths) # Calculate the log-likelihood of the first test sample being generated by this model x, y = test_data[0] model.score(x)
- __init__(*, n_states=5, n_components=3, covariance=CovarianceMode.SPHERICAL, topology=TopologyMode.LEFT_RIGHT, random_state=None, hmmlearn_kwargs=None)¶
Initializes the
GaussianMixtureHMM.- Parameters:
self (GaussianMixtureHMM) –
n_states (int) – Number of states in the Markov chain.
n_components (int) – Number of Gaussian components in the mixture emission distribution for each state.
covariance (CovarianceMode) – Type of covariance matrix in the mixture emission distribution for each state - see Covariance matrix types.
topology (TopologyMode | None) –
Transition topology of the Markov chain — see Topologies.
If
None, behaves the same as'ergodic'but with hmmlearn initialization.random_state (int | RandomState | None) – Seed or
numpy.random.RandomStateobject for reproducible pseudo-randomness.hmmlearn_kwargs (dict[str, Any] | None) – Additional key-word arguments provided to the hmmlearn HMM constructor.
- Return type:
- aic(X, *, lengths=None)¶
The Akaike information criterion of the model, evaluated with the maximum likelihood of
X.- Parameters:
- Returns:
The Akaike information criterion.
- Return type:
Notes
This method requires a trained model — see
fit().
- bic(X, *, lengths=None)¶
The Bayesian information criterion of the model, evaluated with the maximum likelihood of
X.- Parameters:
- Returns:
The Bayesian information criterion.
- Return type:
Notes
This method requires a trained model — see
fit().
- fit(X, *, lengths=None)¶
Fit the HMM to the sequences in
X, using the Baum—Welch algorithm.- Parameters:
- Returns:
The fitted HMM.
- Return type:
BaseHMM
- freeze(params=None, /)¶
Freeze the trainable parameters of the HMM, preventing them from be updated during the Baum—Welch algorithm.
- Parameters:
self (GaussianMixtureHMM) –
params (str | None) –
A string specifying which parameters to freeze. Can contain a combination of:
's'for initial state probabilities,'t'for transition probabilities,'m'for emission distribution means,'c'for emission distribution covariances,'w'for emission distribution mixture weights.
- Return type:
None
See also
unfreezeUnfreeze the trainable parameters of the HMM, allowing them to be updated during the Baum—Welch algorithm.
- score(x, /)¶
Calculate the log-likelihood of the HMM generating a single observation sequence.
- Parameters:
- Returns:
The log-likelihood.
- Return type:
Notes
This method requires a trained model — see
fit().
- set_state_covars(covars, /)¶
Set the covariance matrices of the state emission distributions.
If this method is not called, covariance matrices will be initialized by hmmlearn.
- Parameters:
self (GaussianMixtureHMM) –
covars (ndarray[Any, dtype[float64]]) – Array of covariance values.
- Return type:
None
Notes
If used, this method should normally be called before
fit().
- set_state_means(means, /)¶
Set the mean vectors of the state emission distributions.
If this method is not called, mean vectors will be initialized by hmmlearn.
- Parameters:
self (GaussianMixtureHMM) –
means (ndarray[Any, dtype[float64]]) – Array of mean values.
- Return type:
None
Notes
If used, this method should normally be called before
fit().
- set_state_start_probs(probs=TransitionMode.RANDOM, /)¶
Set the initial state probabilities.
If this method is not called, initial state probabilities are initialized depending on the value of
topologyprovided to__init__().If
topologywas set to'ergodic','left-right'or'linear', then random probabilities will be assigned according to the topology by callingset_state_start_probs()withprobs='random'.If
topologywas set toNone, then initial state probabilities will be initialized by hmmlearn.
- Parameters:
self (BaseHMM) –
probs (ndarray[Any, dtype[float64]] | TransitionMode) –
Probabilities or probability type to assign as initial state probabilities.
If an
Array, should be a vector of starting probabilities for each state.If
'uniform', there is an equal probability of starting in any state.If
'random', the vector of initial state probabilities is sampled from a Dirichlet distribution with unit concentration parameters.
- Return type:
None
Notes
If used, this method should normally be called before
fit().
- set_state_transition_probs(probs=TransitionMode.RANDOM, /)¶
Set the transition probability matrix.
If this method is not called, transition probabilities are initialized depending on the value of
topologyprovided to__init__():If
topologywas set to'ergodic','left-right'or'linear', then random probabilities will be assigned according to the topology by callingset_state_transition_probs()withvalue='random'.If
topologywas set toNone, then initial state probabilities will be initialized by hmmlearn.
- Parameters:
self (BaseHMM) –
probs (ndarray[Any, dtype[float64]] | TransitionMode) –
Probabilities or probability type to assign as state transition probabilities.
If an
Array, should be a matrix of probabilities where each row must some to one and represents the probabilities of transitioning out of a state.If
'uniform', for each state there is an equal probability of transitioning to any state permitted by the topology.If
'random', the vector of transition probabilities for each row is sampled from a Dirichlet distribution with unit concentration parameters, according to the shape of the topology.
- Return type:
None
Notes
If used, this method should normally be called before
fit().
- set_state_weights(weights, /)¶
Set the component mixture weights of the state emission distributions.
If this method is not called, component mixture weights will be initialized by hmmlearn.
- Parameters:
self (GaussianMixtureHMM) –
weights (ndarray[Any, dtype[float64]]) – Array of component mixture weights.
- Return type:
None
Notes
If used, this method should normally be called before
fit().
- unfreeze(params=None, /)¶
Unfreeze the trainable parameters of the HMM, allowing them to be updated during the Baum—Welch algorithm.
- Parameters:
self (GaussianMixtureHMM) –
params (str | None) –
A string specifying which parameters to unfreeze. Can contain a combination of:
's'for initial state probabilities,'t'for transition probabilities,'m'for emission distribution means,'c'for emission distribution covariances,'w'for emission distribution mixture weights.
- Return type:
None
See also
freezeFreeze the trainable parameters of the HMM, preventing them from be updated during the Baum—Welch algorithm.
- covariance: CovarianceMode¶
Type of covariance matrix in the emission model mixture distribution for each state.
- hmmlearn_kwargs: dict[str, Any]¶
Additional key-word arguments provided to the hmmlearn HMM constructor.
- n_components: int¶
Number of Gaussian components in the emission model mixture distribution for each state.
- random_state: int | RandomState | None¶
Seed or
numpy.random.RandomStateobject for reproducible pseudo-randomness.
- topology: TopologyMode¶
Transition topology of the Markov chain — see Topologies.